Fluorescence in situ hybridization (FISH)
This technique was developed in the late 1980s and is a powerful method to detect translocations (rearrangements among chromosomes).
For the development of FISH, it was necessary to isolate each human chromosome. Subsequently, DNA from these chromosomes was fragmented and put into bacterial cells to amplify it (produce many copies). In this way, a large number of copies of DNA from each chromosome can be obtained.
These amplified DNA fragments are labeled with appropriate fluorescent (light-emitting) dyes and allowed to hybridize (attach) to metaphase chromosomes. The fluorescent-labeled DNAs will attach to the analogous chromosomes from which they were derived. (DNA fragments with the same base sequences have the characteristic of attaching to each other.)
In this way, if a part of a painted chromosome (yellow, for example) had undergone an exchange with another, non-painted chromosomes (stained red), it is possible to detect the aberration (termed a reciprocal translocation) because the aberrant chromosome contains both yellow and red segments. Usually, a pair of bi-colored chromosomes can be detected in one metaphase because two chromosomes typically exchange a part of their DNA.
Reciprocal translocations are difficult to detect by simple staining the entire set of chromosomes with a single material, such as with Giemsa. Imagine, for example, a case with which the two exchanged segments had similar lengths. The two translocated chromosomes should appear perfectly normal, both in shape and length. If we employ FISH, however, such translocation can be clearly detected.
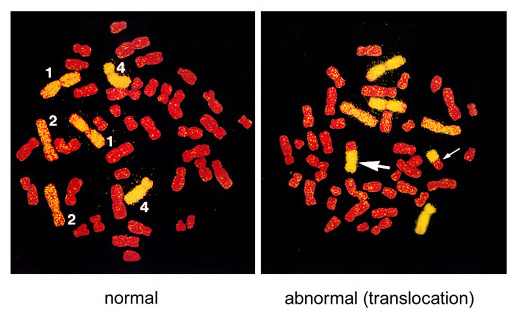
Figure. An example of FISH-treated metaphase chromosomes
Here, chromosomes 1, 2, and 4 were labeled yellow with FISH and the other chromosomes were stained red. Translocations between yellow and red chromosomes are detected. The left picture represents a normal cell (the numbers in the figure indicate chromosome numbers) and the right picture is an example of reciprocal translocation with two bi-color chromosomes (indicated by two arrows).
